Rafflesia Flower
|
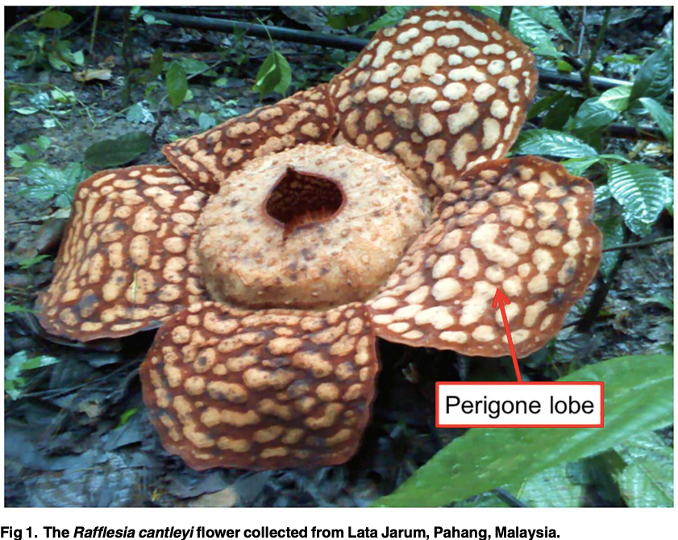
Rafflesia is a biologically enigmatic species that is very rare in occurrence and possesses an extraordinary morphology. It is considered a national treasure of Malaysia.
This parasitic plant produces a gigantic flower up to one metre in diameter with no leaf, stem or root. Little is known about the floral biology of this species, especially at the molecular level. |
Transcriptomics
|
Perigone lobe transcriptome analysis provides insights into Rafflesia cantleyi flower development
We have generated and characterised the transcriptome of the Rafflesia cantleyi flower, and performed a comparison with the transcriptome of its floral bud to predict genes that are expressed and regulated during flower development.
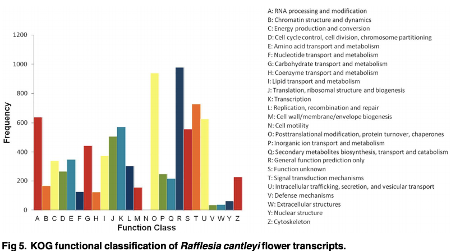
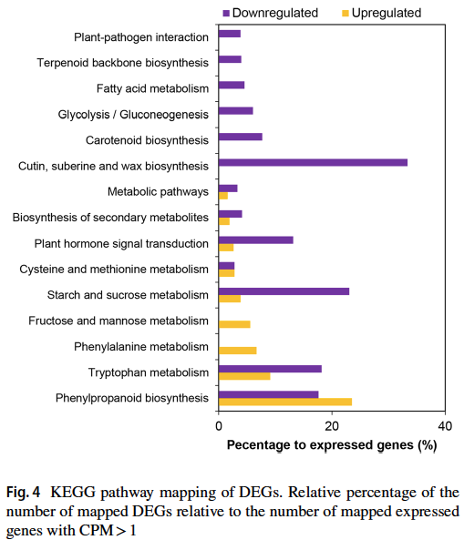
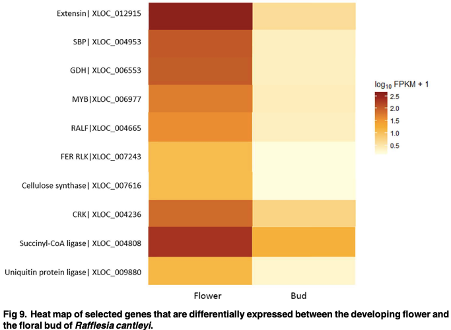
Approximately 40 million sequencing reads were generated and assembled de novo into 18,053 transcripts with an average length of 641 bp. Of these, more than 79% of the transcripts had significant matches to annotated sequences in the public protein database. A total of 11,756 and 7,891 transcripts were assigned to Gene Ontology categories and clusters of orthologous groups, respectively. In addition, 6,019 transcripts could be mapped to 129 pathways in Kyoto Encyclopaedia of Genes and Genomes Pathway database. Digital abundance analysis identified 52 transcripts with very high expression in the flower transcriptome of R. cantleyi. Subsequently, analysis of differential expression between developing flower and the floral bud revealed a set of 105 transcripts with potential role in flower development. |
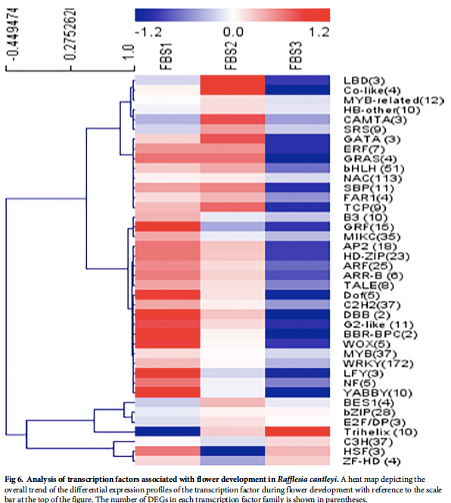
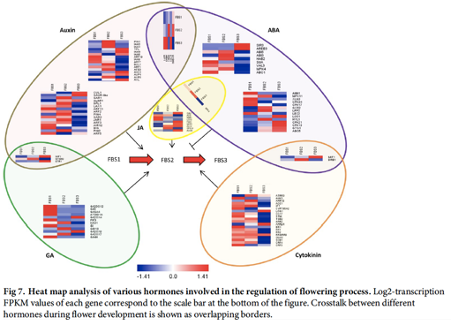
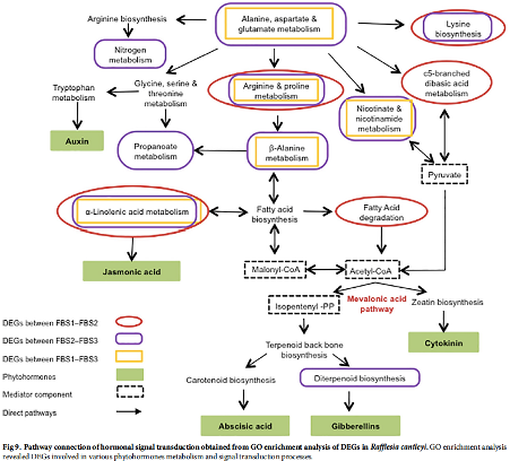
Transcriptome landscape of Rafflesia cantleyi floral buds reveals insights into the roles of transcription factors and phytohormones in flower development
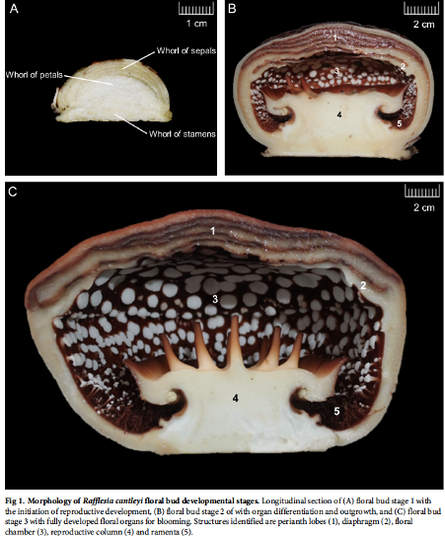
Rafflesia possesses unique biological features and known primarily for producing the world’s largest and existing as a single flower. However, to date, little is known about key regulators participating in Rafflesia flower development. To understand the molecular mechanism that regulates Rafflesia cantleyi flower development, RNA-seq data from three developmental stages of floral bud, representing the floral organ primordia initiation, floral organ differentiation, and floral bud outgrowth, were analysed. A total of 89,890 transcripts were assembled of which up to 35% could be annotated based on homology search. Advanced transcriptome analysis using K-mean clustering on the differentially expressed genes (DEGs) was able to identify 12 expression clusters that reflect major trends and key transitional states, which correlate to specific developmental stages. Comparative gene expression analysis of different floral bud stages identified various transcription factors related to flower development. The members of WRKY, NAC, bHLH, and MYB families are the most represented among the DEGs, suggesting their important functions in flower development. Furthermore, pathway enrichment analysis also revealed DEGs that are involved in various phytohormone signal transduction events such as auxin and auxin transport, cytokinin and gibberellin biosynthesis. |
Significance
Our work presents transcriptome analysis for the developing flower of R. cantleyi. Genes potentially involved in the growth and development of the R. cantleyi flower were identified and provide insights into biological processes that occur during flower development.
Results imply that transcription factors and phytohormone signalling pathways play major role in Rafflesia floral bud development. These studies provide invaluable resources for molecular studies of the flower development process in Rafflesia. The identification of DEGs provides opportunity for further studies to elucidate their functions and contribute towards the understanding of flower development in Rafflesia and other plant species.
Results imply that transcription factors and phytohormone signalling pathways play major role in Rafflesia floral bud development. These studies provide invaluable resources for molecular studies of the flower development process in Rafflesia. The identification of DEGs provides opportunity for further studies to elucidate their functions and contribute towards the understanding of flower development in Rafflesia and other plant species.
References
- Amini S, Rosli K, Abu-Bakar M-F, Alias H, Mat-Isa M-N, Juhari M-A-A, Haji-Adam J, Goh H-H & Wan K-L (2019) Transcriptome landscape of Rafflesia cantleyi floral buds reveals insights into the roles of transcription factors and phytohormones in flower development PLoS One 14 (12): e0226338.
- Amini, S., Alias, H., Aizat-Juhari, M. A., Mat-Isa, M. N., Adam, J. H., Goh, H.H. & Wan, K. L. (2017) RNA-Seq Data from Different Developmental Stages of Rafflesia cantleyi Floral Buds. Genomics Data 14:5-6.
- Lee X-W, Mat-Isa M-N, Mohd-Elias N-A, Aizat-Juhari MA, Goh H-H, Dear PH, Chow K-S, Haji Adam J, Mohamed R, Firdaus-Raih M & Wan K-L. (2016) Perigone lobe transcriptome analysis provides insights into Rafflesia cantleyi flower development. PloS ONE, 11(12), e0167958, doi:10.1371/journal.pone.0167958
- GENERATION AND CHARACTERIZATION OF THE RAFFLESIA GENOME SEQUENCE Research University Grant: LAUREATE-2013-001 [1 Jan 2014 – 31 Dec 2016] – Co-researcher
Copyright © 2022 UKM